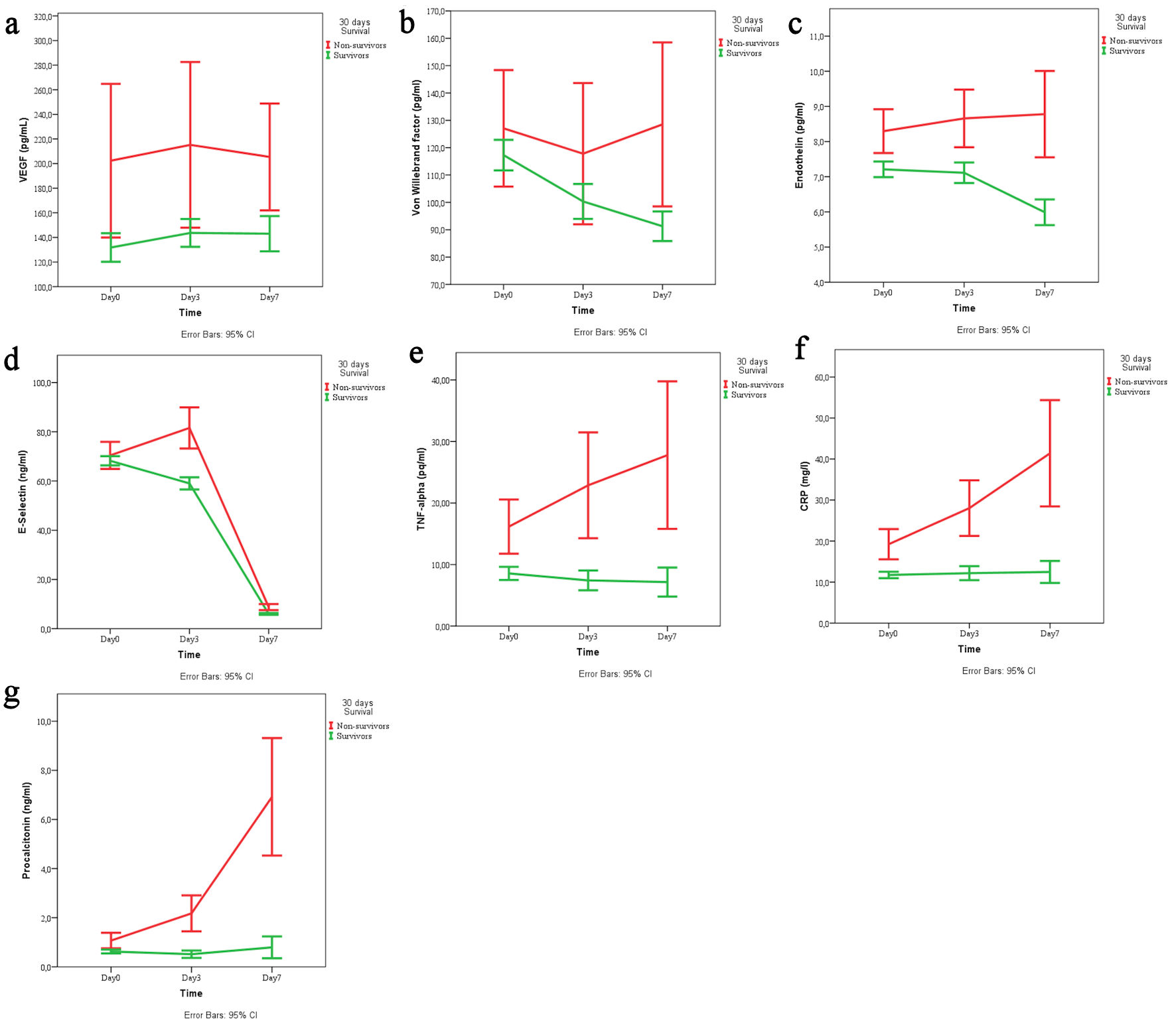
↓ Figure 1. Temporal trends of (a) VEGF, (b) von Willebrand factor, (c) endothelin, (d) E-selectin, (e) TNF-α, (f) CRP, and (g) procalcitonin across the three time points (day 1, 3, 7) stratified by survival status (survivors vs. non-survivors). VEGF: vascular endothelial growth factor; TNF-α: tumor necrosis factor-α; CRP: C-reactive protein; CI: confidence interval.
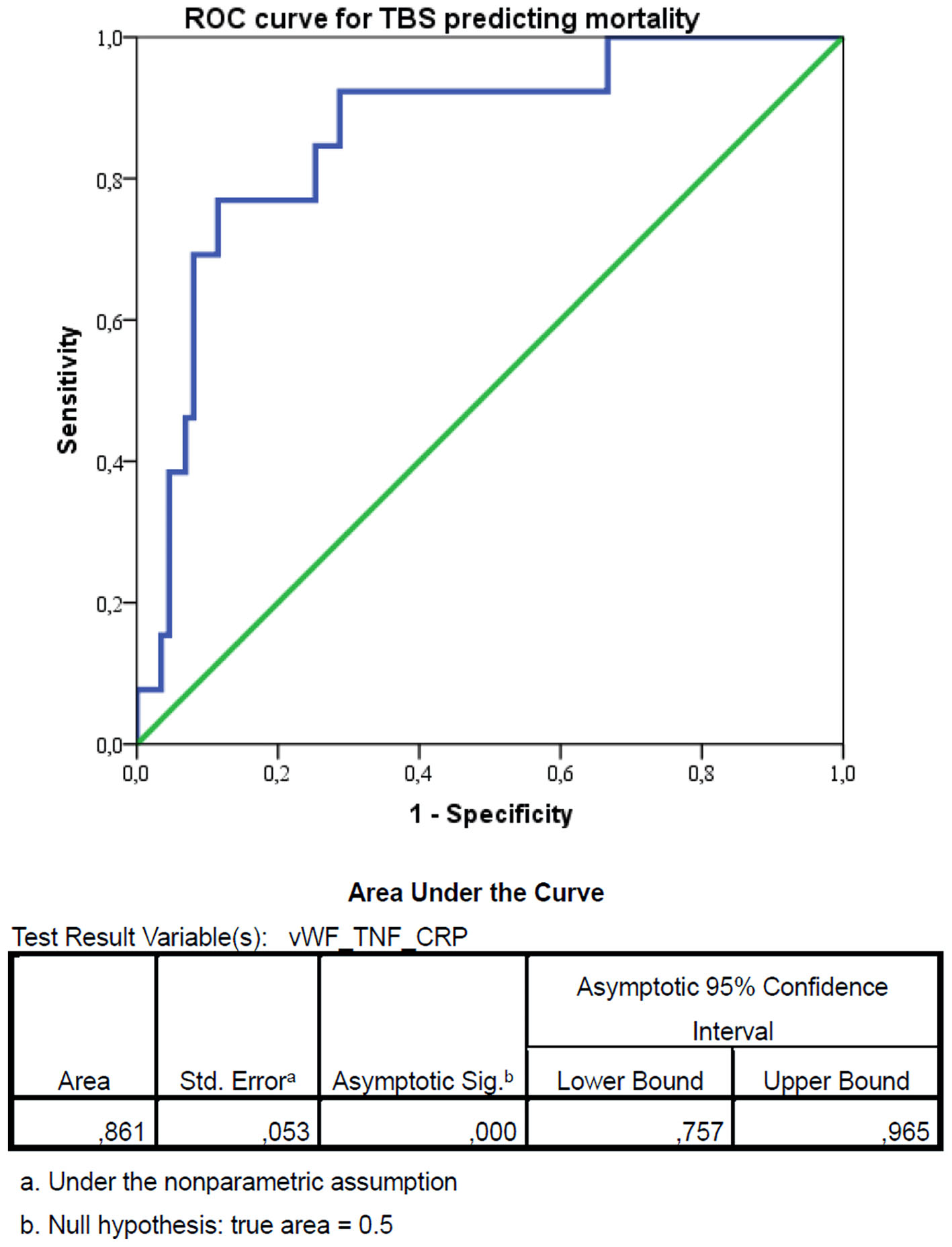
↓ Figure 3. Receiver operating characteristic (ROC) curve for the triple biomarker score (TBS) predicting in-hospital mortality. The optimal cut-off value was identified as 4952.65, using Youden’s index maximization, which yielded a sensitivity of 92.3% and a specificity of 69.0% (Youden’s index J = 0.613). vWF: von Willebrand factor; TNF-α: tumor necrosis factor-α; CRP: C-reactive protein; CI: confidence interval.
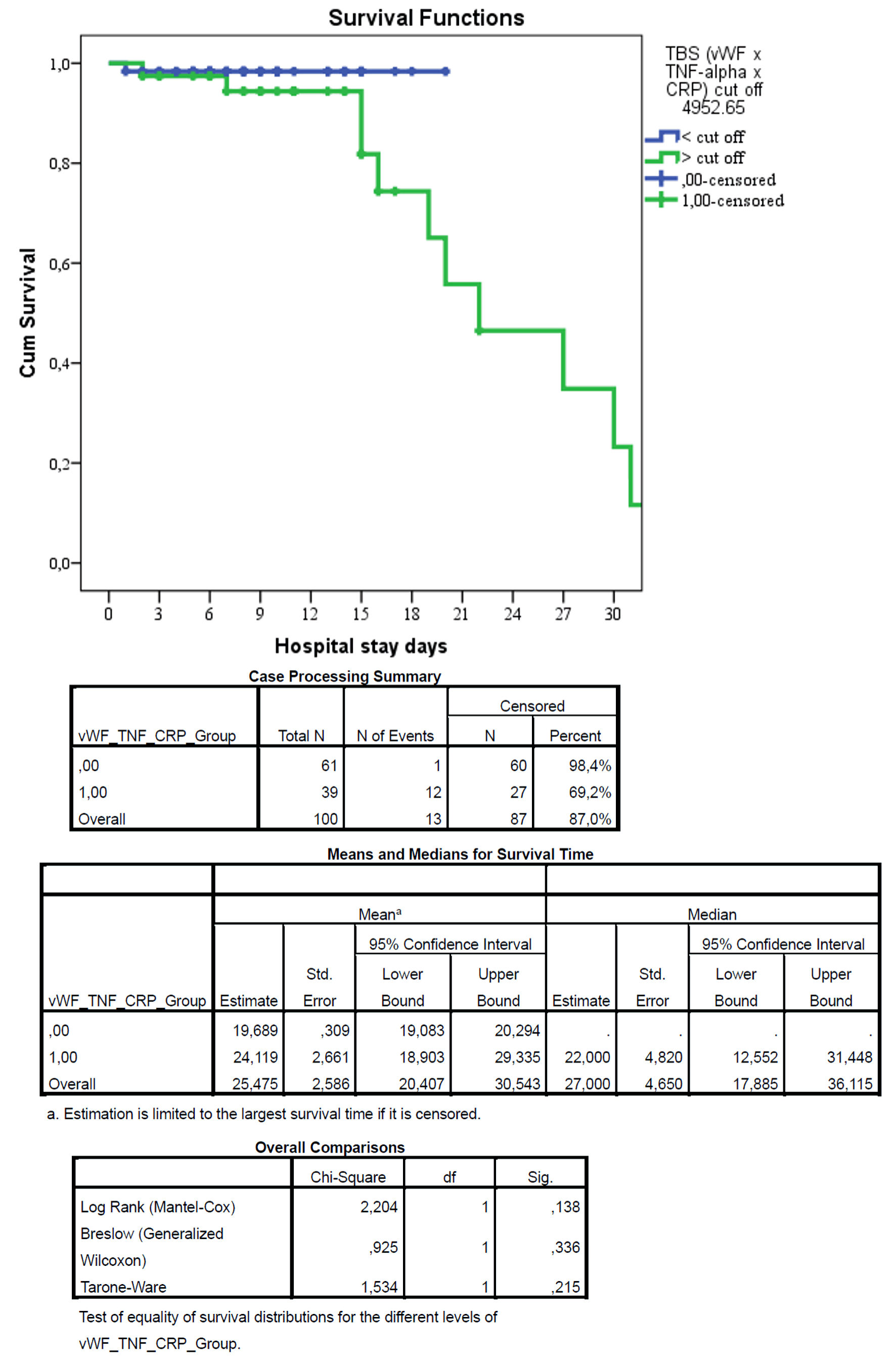
↓ Figure 4. Kaplan–Meier survival curves for patients stratified by triple biomarker score (TBS). Kaplan–Meier survival analysis demonstrated no statistically significant difference between groups (log-rank test: χ2 = 2.204, P = 0.138). Although mortality was numerically lower in group 0 compared to group 1, the difference did not reach statistical significance, likely due to the limited number of events. Censored patients are indicated by tick marks. The low TBS group had significantly higher survival (98.4%) compared to the high TBS group (69.2%), with a log-rank P value < 0.001. The table below shows the number of patients at risk at each time point.